Ziqing Liu, Ph.D.
Research Associate (2018 – present)
Post-Doctoral Fellow (2014 – 2017)
University of North Carolina-Chapel Hill – Department of Pathology and Laboratory Medicine
Principal Investigator: Dr. Li Qian; Study area: direct cardiac reprogramming
Contact Info: ziqing@email.unc.edu
Education:
Indiana University (IU), Indianapolis, IN – Ph. D. in Microbiology and Immunology (2014)
University of Science and Technology of China (USTC), Hefei, China – B.S. in Life Science (2008)
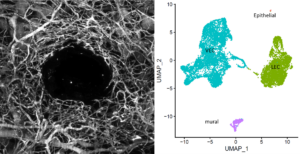
Research: Understanding cellular and molecular heterogeneity of endothelial cells in blood vessel formation, homeostasis, and wound healing by utilization of single-cell RNA sequencing and a variety of in vitro cell culture and in vivo animal models.
Publications:
Liu Z, Ruter DL, Quigley KM, Tanke N, Jiang Y, Bautch VL (2021). Single-Cell RNA Sequencing Reveals Endothelial Cell Transcriptome Heterogeneity under Homeostatic Laminar Flow. Arteriosclerosis, Thrombosis, and Vascular Biology. 2021 Aug 26:ATVBAHA121316797. Featured article of the issue.
Ruter DL, Liu Z, Ngo KM, X S, Marvin A, Buglak DB, Kidder EJ, Bautch VL (2021). SMAD6 Integrates Endothelial Cell Homeostatic Flow Responses Downstream of Notch. Angiogenesis. 2021 May;24(2):387-398.
Ma H*, Liu Z*, Yang Y*, Feng D, Dong Y, Garbutt T, Hu Z, Wang L, Luan H, Cooper C, Li Y, Welch J, Qian L, Liu J (2021). Functional Coordination of Non-myocytes Plays a Key Role in Adult Zebrafish Heart Regeneration. EMBO Reports. 2021 e52901 https://doi.org/10.15252/embr.202152901. *co-first author.
Wang L, Ma H, Huang P, Xie Y, Near D, Wang H, Xu J, Yang Y, Xu Y, Garbutt T, Zhou Y, Liu Z, Yin C, Bressan M, Taylor JM, Liu JD and Qian L (2020). Downregulation of Beclin1 Promotes Direct Cardiac Reprogramming. Sci Transl Med 12(566): eaay7856.
Zhou Y*, Liu Z*, Welch JD, Gao X, Wang L, Garbutt T, Keepers B, Ma H, Prins JF, Shen WN,
Liu JD, Qian L (2019). Single-Cell Transcriptomic Analyses of Cell Fate Transitions during Human Cardiac Reprogramming. Cell Stem Cell 25 (1), 149-164. *co-first author.
Ma H, Yu S, Liu X, Zhang Y, Fakadej T, Liu Z, Yin CY, Shen WN, Locasale JW, Taylor JM, Qian L, Liu JD (2019). Lin28a Regulates Pathological Cardiac Hypertrophic Growth through Pck2-Mediated Enhancement of Anabolic Synthesis. Circulation 139 (14), 1725-1740.
Zhou Y, Alimohamadi S, Wang L, Liu Z, Wall JB, Yin C, Liu J, Qian L (2018). A Loss of Function Screen of Epigenetic Modifiers and Splicing Factors during Early Stage of Cardiac Reprogramming. Stem cells international.
Liu Z*, Wang L*, Welch JD*, Ma H, Zhou Y, Vaseghi HR, Yu S, Wall JB, Alimohamadi S, Zheng M, Yin CY, Shen WN, Prins JF, Liu JD, Qian L (2017). Single cell transcriptomics reconstructs lineage conversion from fibroblast to cardiomyocyte. Nature, 551, 100-104 *: co-first author.
Zhou Y, Wang L, Liu Z, Alimohamadi S, Yin CY, Liu JD, Qian L (2017). Comparative gene expression analyses reveal distinct molecular signature between induced cardiomyocytes and induced pluripotent stem cell-derived cardiomyocytes. Cell Report, 20(13), 3014-3024.
Liu Z, Chen O, Wall JB, Zheng M, Zhou Y, Wang L, Vaseghi HR, Qian L, Liu JD (2017). Systematic comparison of 2A peptides for cloning multi-genes in a polycistronic vector. Scientific Reports, 7: 2193.
Liu Z, Chen O, Zheng M, Wang L, Zhou Y, Yin CY, Liu JD, Qian L (2016). Re-patterning of H3K27me3, H3K4me3 and DNA methylation during fibroblast conversion into induced cardiomyocytes. Stem Cell Research. 16 (2), 507–518.
Zhou Y, Wang L, Vaseghi HR, Liu Z, Lu R, Alimohamadi S, Yin CY, Fu JD, Wang GG, Liu JD, Qian L (2016). Bmi1 Functions as a Critical Epigenetic Modulator during Early Phase of Direct Cardiac Reprogramming. Cell Stem Cell, 18 (3), 382-95. Commentary: Herrer D and Benard A. Bmi1-mediated epigenetic signature acts as a critical barrier for direct reprogramming to mature cardiomyocytes. Stem Cell Investigation. 2016 Jul 20; 3-28.
Wang L, Liu Z, Yin C, Zhou Y, Liu J, Qian L (2015). Improved Generation of Induced Cardiomyocytes Using a Polycistronic Construct Expressing Optimal Ratio of Gata4, Mef2c and Tbx5. JoVE (Journal of Visualized Experiments) e53426-e53426.
Wang L*, Liu Z*, Yin C, Asfour H, Chen O, Li Y, Bursac N, Liu J, Qian L (2014). Stoichiometry of Gata4, Mef2c, and Tbx5 influences the efficiency and quality of induced cardiac myocyte reprogramming. Circulation Research, 116 (2), 237-244. *: co-first author. Editorial’s pick of the issue. Editorial: Muraoka N, Ieda M. Stoichiometry of transcription factors is critical for cardiac reprogramming. Circulation Research. 2015 Jan 16; 116: 216-218.
Liu Z, F Zhao, He JJ (2014). Hepatitis C virus (HCV) interaction with astrocytes: non-productive infection and induction of IL-18. Journal of Neurovirology, 20 (3), 278-293
Liu Z, X Zhang, Q Yu, He JJ (2014). Exosome-associated hepatitis C virus in cell cultures and patient plasma. Biochemical and biophysical research communications, 455 (3), 218-222
Liu Z, He JJ (2013). Cell-cell contact-mediated hepatitis C virus (HCV) transfer, productive infection and replication and its requirement for HCV receptors. Journal of Virology, 87(15):8545-58.
Park IW, Ndjomou J, Wen Y, Liu Z, Ridgway ND, Kao CC, He JJ (2013). Inhibition of HCV Replication by Oxysterol-Binding Protein-Related Protein 4 (ORP4) through Interaction with HCV NS5B and Alteration of Lipid Droplet Formation. Plos One, 8(9):e75648.
Amet T, Ghabril M, Chalasani N, Byrd D, Hu N, Grantham A, Liu Z, Qin X, He JJ, Yu Q (2012). CD59 incorporation protects hepatitis C virus against complement-mediated destruction. Hepatology, 55(2):354-63.
Zhan J, Ding B, Ma R, Ma X, Su X, Zhao Y, Liu Z, Wu J, Liu H (2010). Develop reusable and combinable designs for transcriptional logic gates. Molecular Systems Biology, 6:388.


